Coronavirus variants: how did they evolve and what do they mean?
Professor of Microbial Evolution at The Milner Centre for Evolution, University of Bath
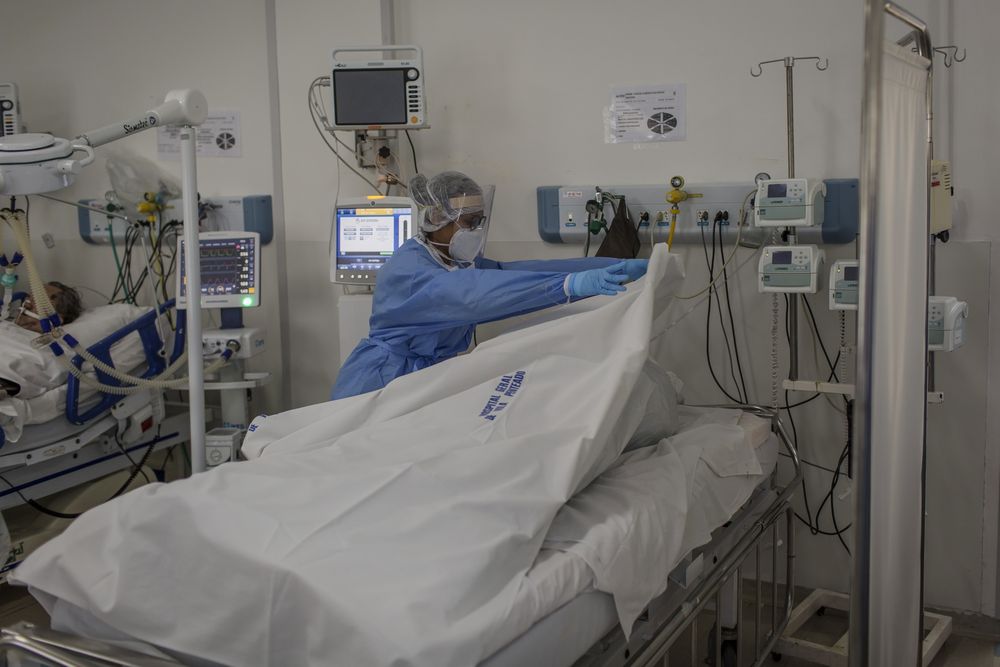
When supplies of oxygen at hospitals in Manaus, Brazil, recently ran out, the airforce was called in for emergency evacuations while healthcare workers frantically tried to save lives with manual ventilation. For those that could not be saved, there was only morphine and a final hand-squeeze.
As calamitous as the situation is for those affected, the devastating surge in COVID cases in Manaus over the last few weeks has set alarm bells ringing ever more loudly for governments and agencies around the world struggling to manage the pandemic. Cases continue to surge in the UK and South Africa and, as in Manaus, they appear to be mainly due to the emergence of new variants of the coronavirus.
Three variants causing concern
The naming of these “variants of concern” – as scientists refer to them – is somewhat muddled. For simplicity, they are referred to here as the Brazilian, South African and UK variant. All have emerged recently, and all have picked up several mutations that mark a distinct shift in the evolution of the virus. Similar variants are almost certainly out there spreading under the radar. More are likely to evolve.
Despite arising independently on three different continents, the three variants share striking similarities. Each has picked up several mutations over a short time, with many in the gene providing the instructions to make the virus’s spike protein.
The spike protein is where the key battles between human and virus are being waged, including the vaccines. It is the key to how the virus interacts with the human body, both regarding the immune response and in binding to and entering human airway cells.
Not only have several mutations affected this protein, but identical mutations have cropped up independently both in the variants of concern and in other viral lineages. In effect, the virus has repeatedly stumbled across the same evolutionary solutions to specific challenges. This phenomenon is known as evolutionary convergence (consider the independent evolution of wings in bats, birds and insects).
Understanding how these mutations might affect the coronavirus’s behaviour at the molecular level is difficult. Work to bridge the gap between each variant’s “genotype” (the mutations) and its “phenotype” (how quickly it spreads) is being ramped up in the UK and elsewhere, but will require a sustained multidisciplinary effort.
Constellations of mutations
The task is made more difficult because several mutations have accumulated in these variants (so-called constellations). The UK variant, for example, has 23 separate mutations, representing a remarkable evolutionary jump with no known intermediate variants (like there are “missing links” in the evolutionary chain).
Although not all the mutations are thought to be important, the effect of any individual mutation might be changed by the presence of other mutations (an effect called epistasis). This greatly complicates the problem of figuring out precisely what these mutations are doing and of assessing the risk of newly emerging variants from the sequence data alone.
Despite these complexities, a combination of computational analysis and laboratory experiments have yielded valuable evidence of the effect of these mutations. For example, one mutation found in all three variants is N501Y. The name refers to an alteration in the spike protein, where the type of amino-acid molecule located in position 501 has changed from asparagine (N) to tyrosine (Y).
Position 501 is on the receptor-binding domain – part of the spike protein that attaches to a particular receptor (ACE2) on cells in the human body – and this change appears to strengthen the binding between the virus and human cells. Yet for reasons that remain unclear, the effect of N501Y is greatly amplified when combined with other mutations.
Other mutations in the spike protein offer the virus some protection from the immune response. Examples include E484K (found in the Brazilian and South African variants, but not the UK variant), and a mutation in the UK variant in which two amino acids are deleted (del69-70) and which is repeatedly found in combination with mutations in the receptor-binding domain.
Selection pressure
Specific evolutionary challenges and selection pressures that favour the survival of some variants of the virus over others may be driving the emergence of the variants of concern. This would help to explain why they acquire several mutations so quickly, or why these variants are starting to emerge now.
A plausible explanation for the emergence of the UK variant is that it arose in a single chronically infected person with a weakened immune system who was being treated with convalescent plasma (antibodies from a recovered patient). This would have given a strong advantage to any variant that could resist the therapeutic antibodies. But it remains a theory.
A second possibility relates to the emergence of the Brazilian variant. The current wave of infection in Manaus is only the latest COVID disaster to hit this city. Previous waves may have led to 76% of the population being infected. The resulting high level of immunity in the population may have given an advantage to mutations in the spike protein.
Although these variants are causing concern, we should remain confident that the vaccines will ultimately prove successful in ending the pandemic and allow a return to normality. There is currently no evidence that the vaccines are less effective against the new variants. While it remains impossible to be certain whether, or how, the virus will make further evolutionary jumps when confronted by the vaccines, modifications to vaccine design should ensure that we stay one step ahead. ( From : The Conversation )

















Facebook Comments